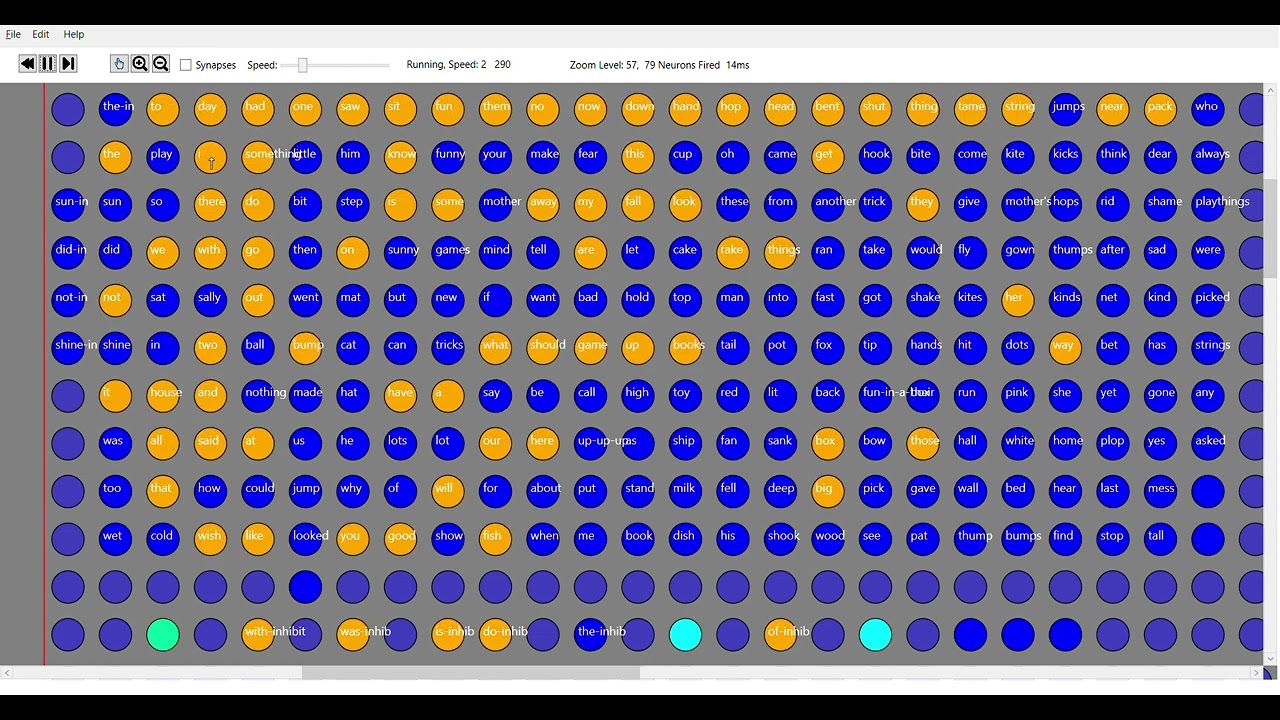
The issue of coding longer sequences in an SDR has come up in my Chaos and sequential AI models thread with @complyue . That we need to represent letters with SDR, so we can capture code for paths back along a sequence:
I recall some work done by Felix Andrews @floybix on this, and discussed on the predecessor to this forum, back, initially together with me, starting in 2014. As I recall, basically, making sequence predicting links from only a random selection of nodes in an SDR for a state, could distinguish a path, back, before that state, and onwards, so that the path through that state would be distinguishable in subsequent states too.
This should be important, because the immediate task we are looking at is to represent words as sequences of letters. So we want to have the code for a letter like “e”, in a word like “place”, also be distinct according to the path which arrived at it, through “p”, “l”, “a”, “c”. So, in a way, the SDR for “e” in this context can be distinct, although sharing enough commonality with “e” arrived at in paths through other words, that it can still be identified as a state for “e”.
Firstly, is this something still in HTM? Or if not is there an archive online anywhere for the pre-HTM Formum, NuPIC discussion list where it was discussed?
I’ve been able to find a few links to Felix’s work online. It mostly centered around his port of HTM to Clojure, called Comportex. I see there is still a github repository:
There’s the document on Felix’s continuation of our sequence experiments:
http://viewer.gorilla-repl.org/view.html?source=gist&id=95da4401dc7293e02df3&filename=seq-replay.clj
I think this short demo is of the same (in 2016):
HTM higher-order sequence learning - SDRs
There’s also this presentation by Felix at the 2014 fall hackathon. Though it may just be for the Clojure port generally:
HTM in Clojure [DEMO #6] (2014 Fall NuPIC Hackathon)
I recall Felix was able to demonstrate the recall of arbitrarily long sequences. It would be good to recover some of those old insights. Or perhaps the current theory of HTM has superseded it??
I don’t suppose @floybix still monitors this forum!

![HTM in Clojure [DEMO #6] (2014 Fall NuPIC Hackathon)](https://img.youtube.com/vi/hE6alw_HHrk/maxresdefault.jpg)